Compare outcomes from differential analysis based on different imputation methods#
load scores based on
10_1_ald_diff_analysis
Parameters#
Default and set parameters for the notebook.
folder_experiment = 'runs/appl_ald_data/plasma/proteinGroups'
target = 'kleiner'
model_key = 'VAE'
baseline = 'RSN'
out_folder = 'diff_analysis'
selected_statistics = ['p-unc', '-Log10 pvalue', 'qvalue', 'rejected']
disease_ontology = 5082 # code from https://disease-ontology.org/
# split diseases notebook? Query gene names for proteins in file from uniprot?
annotaitons_gene_col = 'PG.Genes'
# Parameters
disease_ontology = 10652
folder_experiment = "runs/alzheimer_study"
target = "AD"
baseline = "PI"
model_key = "DAE"
out_folder = "diff_analysis"
annotaitons_gene_col = "None"
Add set parameters to configuration
root - INFO Removed from global namespace: folder_experiment
root - INFO Removed from global namespace: target
root - INFO Removed from global namespace: model_key
root - INFO Removed from global namespace: baseline
root - INFO Removed from global namespace: out_folder
root - INFO Removed from global namespace: selected_statistics
root - INFO Removed from global namespace: disease_ontology
root - INFO Removed from global namespace: annotaitons_gene_col
root - INFO Already set attribute: folder_experiment has value runs/alzheimer_study
root - INFO Already set attribute: out_folder has value diff_analysis
{'annotaitons_gene_col': 'None',
'baseline': 'PI',
'data': PosixPath('runs/alzheimer_study/data'),
'disease_ontology': 10652,
'folder_experiment': PosixPath('runs/alzheimer_study'),
'freq_features_observed': PosixPath('runs/alzheimer_study/freq_features_observed.csv'),
'model_key': 'DAE',
'out_figures': PosixPath('runs/alzheimer_study/figures'),
'out_folder': PosixPath('runs/alzheimer_study/diff_analysis/AD/PI_vs_DAE'),
'out_metrics': PosixPath('runs/alzheimer_study'),
'out_models': PosixPath('runs/alzheimer_study'),
'out_preds': PosixPath('runs/alzheimer_study/preds'),
'scores_folder': PosixPath('runs/alzheimer_study/diff_analysis/AD/scores'),
'selected_statistics': ['p-unc', '-Log10 pvalue', 'qvalue', 'rejected'],
'target': 'AD'}
Excel file for exports#
files_out = dict()
writer_args = dict(float_format='%.3f')
fname = args.out_folder / 'diff_analysis_compare_methods.xlsx'
files_out[fname.name] = fname
writer = pd.ExcelWriter(fname)
logger.info("Writing to excel file: %s", fname)
root - INFO Writing to excel file: runs/alzheimer_study/diff_analysis/AD/PI_vs_DAE/diff_analysis_compare_methods.xlsx
Load scores#
Load baseline model scores#
Show all statistics, later use selected statistics
| model | PI | ||||||||
|---|---|---|---|---|---|---|---|---|---|
| var | SS | DF | F | p-unc | np2 | -Log10 pvalue | qvalue | rejected | |
| protein groups | Source | ||||||||
| A0A024QZX5;A0A087X1N8;P35237 | AD | 0.261 | 1 | 0.513 | 0.475 | 0.003 | 0.324 | 0.628 | False |
| age | 0.017 | 1 | 0.034 | 0.854 | 0.000 | 0.069 | 0.911 | False | |
| Kiel | 1.577 | 1 | 3.106 | 0.080 | 0.016 | 1.099 | 0.175 | False | |
| Magdeburg | 4.866 | 1 | 9.584 | 0.002 | 0.048 | 2.646 | 0.010 | True | |
| Sweden | 8.181 | 1 | 16.112 | 0.000 | 0.078 | 4.067 | 0.001 | True | |
| ... | ... | ... | ... | ... | ... | ... | ... | ... | ... |
| S4R3U6 | AD | 0.248 | 1 | 0.255 | 0.614 | 0.001 | 0.212 | 0.741 | False |
| age | 0.492 | 1 | 0.505 | 0.478 | 0.003 | 0.320 | 0.631 | False | |
| Kiel | 0.000 | 1 | 0.000 | 0.984 | 0.000 | 0.007 | 0.991 | False | |
| Magdeburg | 1.226 | 1 | 1.259 | 0.263 | 0.007 | 0.580 | 0.421 | False | |
| Sweden | 14.711 | 1 | 15.099 | 0.000 | 0.073 | 3.852 | 0.001 | True | |
7105 rows × 8 columns
Load selected comparison model scores#
| model | DAE | ||||||||
|---|---|---|---|---|---|---|---|---|---|
| var | SS | DF | F | p-unc | np2 | -Log10 pvalue | qvalue | rejected | |
| protein groups | Source | ||||||||
| A0A024QZX5;A0A087X1N8;P35237 | AD | 1.066 | 1 | 7.727 | 0.006 | 0.039 | 2.223 | 0.017 | True |
| age | 0.003 | 1 | 0.021 | 0.886 | 0.000 | 0.053 | 0.926 | False | |
| Kiel | 0.260 | 1 | 1.883 | 0.172 | 0.010 | 0.765 | 0.279 | False | |
| Magdeburg | 0.492 | 1 | 3.565 | 0.061 | 0.018 | 1.218 | 0.121 | False | |
| Sweden | 1.740 | 1 | 12.620 | 0.000 | 0.062 | 3.318 | 0.002 | True | |
| ... | ... | ... | ... | ... | ... | ... | ... | ... | ... |
| S4R3U6 | AD | 2.843 | 1 | 4.846 | 0.029 | 0.025 | 1.539 | 0.066 | False |
| age | 0.601 | 1 | 1.025 | 0.313 | 0.005 | 0.505 | 0.442 | False | |
| Kiel | 3.475 | 1 | 5.924 | 0.016 | 0.030 | 1.800 | 0.039 | True | |
| Magdeburg | 3.375 | 1 | 5.754 | 0.017 | 0.029 | 1.759 | 0.043 | True | |
| Sweden | 15.553 | 1 | 26.514 | 0.000 | 0.122 | 6.188 | 0.000 | True | |
7105 rows × 8 columns
Combined scores#
show only selected statistics for comparsion
| model | DAE | PI | |||||||
|---|---|---|---|---|---|---|---|---|---|
| var | p-unc | -Log10 pvalue | qvalue | rejected | p-unc | -Log10 pvalue | qvalue | rejected | |
| protein groups | Source | ||||||||
| A0A024QZX5;A0A087X1N8;P35237 | AD | 0.006 | 2.223 | 0.017 | True | 0.475 | 0.324 | 0.628 | False |
| Kiel | 0.172 | 0.765 | 0.279 | False | 0.080 | 1.099 | 0.175 | False | |
| Magdeburg | 0.061 | 1.218 | 0.121 | False | 0.002 | 2.646 | 0.010 | True | |
| Sweden | 0.000 | 3.318 | 0.002 | True | 0.000 | 4.067 | 0.001 | True | |
| age | 0.886 | 0.053 | 0.926 | False | 0.854 | 0.069 | 0.911 | False | |
| ... | ... | ... | ... | ... | ... | ... | ... | ... | ... |
| S4R3U6 | AD | 0.029 | 1.539 | 0.066 | False | 0.614 | 0.212 | 0.741 | False |
| Kiel | 0.016 | 1.800 | 0.039 | True | 0.984 | 0.007 | 0.991 | False | |
| Magdeburg | 0.017 | 1.759 | 0.043 | True | 0.263 | 0.580 | 0.421 | False | |
| Sweden | 0.000 | 6.188 | 0.000 | True | 0.000 | 3.852 | 0.001 | True | |
| age | 0.313 | 0.505 | 0.442 | False | 0.478 | 0.320 | 0.631 | False | |
7105 rows × 8 columns
Models in comparison (name mapping)
{'DAE': 'DAE', 'PI': 'PI'}
Describe scores#
| model | DAE | PI | ||||
|---|---|---|---|---|---|---|
| var | p-unc | -Log10 pvalue | qvalue | p-unc | -Log10 pvalue | qvalue |
| count | 7,105.000 | 7,105.000 | 7,105.000 | 7,105.000 | 7,105.000 | 7,105.000 |
| mean | 0.224 | 3.317 | 0.278 | 0.259 | 2.482 | 0.335 |
| std | 0.293 | 6.333 | 0.319 | 0.301 | 5.314 | 0.329 |
| min | 0.000 | 0.000 | 0.000 | 0.000 | 0.001 | 0.000 |
| 25% | 0.001 | 0.408 | 0.002 | 0.004 | 0.334 | 0.015 |
| 50% | 0.061 | 1.216 | 0.122 | 0.119 | 0.925 | 0.238 |
| 75% | 0.391 | 3.275 | 0.521 | 0.463 | 2.435 | 0.618 |
| max | 0.999 | 86.886 | 0.999 | 0.999 | 147.300 | 0.999 |
One to one comparison of by feature:#
/tmp/ipykernel_99922/3761369923.py:2: FutureWarning: Starting with pandas version 3.0 all arguments of to_excel except for the argument 'excel_writer' will be keyword-only.
scores.to_excel(writer, 'scores', **writer_args)
| model | DAE | PI | |||||||
|---|---|---|---|---|---|---|---|---|---|
| var | p-unc | -Log10 pvalue | qvalue | rejected | p-unc | -Log10 pvalue | qvalue | rejected | |
| protein groups | Source | ||||||||
| A0A024QZX5;A0A087X1N8;P35237 | AD | 0.006 | 2.223 | 0.017 | True | 0.475 | 0.324 | 0.628 | False |
| A0A024R0T9;K7ER74;P02655 | AD | 0.034 | 1.473 | 0.075 | False | 0.029 | 1.531 | 0.080 | False |
| A0A024R3W6;A0A024R412;O60462;O60462-2;O60462-3;O60462-4;O60462-5;Q7LBX6;X5D2Q8 | AD | 0.225 | 0.648 | 0.344 | False | 0.089 | 1.050 | 0.191 | False |
| A0A024R644;A0A0A0MRU5;A0A1B0GWI2;O75503 | AD | 0.257 | 0.590 | 0.382 | False | 0.508 | 0.294 | 0.656 | False |
| A0A075B6H7 | AD | 0.019 | 1.728 | 0.046 | True | 0.122 | 0.913 | 0.243 | False |
| ... | ... | ... | ... | ... | ... | ... | ... | ... | ... |
| Q9Y6R7 | AD | 0.175 | 0.756 | 0.284 | False | 0.175 | 0.756 | 0.316 | False |
| Q9Y6X5 | AD | 0.281 | 0.551 | 0.410 | False | 0.044 | 1.354 | 0.110 | False |
| Q9Y6Y8;Q9Y6Y8-2 | AD | 0.083 | 1.079 | 0.157 | False | 0.083 | 1.079 | 0.182 | False |
| Q9Y6Y9 | AD | 0.701 | 0.154 | 0.791 | False | 0.094 | 1.025 | 0.200 | False |
| S4R3U6 | AD | 0.029 | 1.539 | 0.066 | False | 0.614 | 0.212 | 0.741 | False |
1421 rows × 8 columns
And the descriptive statistics of the numeric values:
| model | DAE | PI | ||||
|---|---|---|---|---|---|---|
| var | p-unc | -Log10 pvalue | qvalue | p-unc | -Log10 pvalue | qvalue |
| count | 1,421.000 | 1,421.000 | 1,421.000 | 1,421.000 | 1,421.000 | 1,421.000 |
| mean | 0.237 | 1.613 | 0.297 | 0.257 | 1.407 | 0.337 |
| std | 0.290 | 1.891 | 0.314 | 0.296 | 1.627 | 0.320 |
| min | 0.000 | 0.001 | 0.000 | 0.000 | 0.001 | 0.000 |
| 25% | 0.006 | 0.383 | 0.018 | 0.012 | 0.342 | 0.040 |
| 50% | 0.084 | 1.074 | 0.158 | 0.119 | 0.925 | 0.238 |
| 75% | 0.414 | 2.209 | 0.544 | 0.455 | 1.907 | 0.610 |
| max | 0.997 | 24.290 | 0.997 | 0.997 | 21.214 | 0.998 |
and the boolean decision values
| model | DAE | PI |
|---|---|---|
| var | rejected | rejected |
| count | 1421 | 1421 |
| unique | 2 | 2 |
| top | False | False |
| freq | 933 | 1033 |
Load frequencies of observed features#
| data | |
|---|---|
| frequency | |
| protein groups | |
| A0A024QZX5;A0A087X1N8;P35237 | 186 |
| A0A024R0T9;K7ER74;P02655 | 195 |
| A0A024R3W6;A0A024R412;O60462;O60462-2;O60462-3;O60462-4;O60462-5;Q7LBX6;X5D2Q8 | 174 |
| A0A024R644;A0A0A0MRU5;A0A1B0GWI2;O75503 | 196 |
| A0A075B6H7 | 91 |
| ... | ... |
| Q9Y6R7 | 197 |
| Q9Y6X5 | 173 |
| Q9Y6Y8;Q9Y6Y8-2 | 197 |
| Q9Y6Y9 | 119 |
| S4R3U6 | 126 |
1421 rows × 1 columns
Plot qvalues of both models with annotated decisions#
Prepare data for plotting (qvalues)
| DAE | PI | frequency | Differential Analysis Comparison | |
|---|---|---|---|---|
| protein groups | ||||
| A0A024QZX5;A0A087X1N8;P35237 | 0.017 | 0.628 | 186 | DAE (yes) - PI (no) |
| A0A024R0T9;K7ER74;P02655 | 0.075 | 0.080 | 195 | DAE (no) - PI (no) |
| A0A024R3W6;A0A024R412;O60462;O60462-2;O60462-3;O60462-4;O60462-5;Q7LBX6;X5D2Q8 | 0.344 | 0.191 | 174 | DAE (no) - PI (no) |
| A0A024R644;A0A0A0MRU5;A0A1B0GWI2;O75503 | 0.382 | 0.656 | 196 | DAE (no) - PI (no) |
| A0A075B6H7 | 0.046 | 0.243 | 91 | DAE (yes) - PI (no) |
| ... | ... | ... | ... | ... |
| Q9Y6R7 | 0.284 | 0.316 | 197 | DAE (no) - PI (no) |
| Q9Y6X5 | 0.410 | 0.110 | 173 | DAE (no) - PI (no) |
| Q9Y6Y8;Q9Y6Y8-2 | 0.157 | 0.182 | 197 | DAE (no) - PI (no) |
| Q9Y6Y9 | 0.791 | 0.200 | 119 | DAE (no) - PI (no) |
| S4R3U6 | 0.066 | 0.741 | 126 | DAE (no) - PI (no) |
1421 rows × 4 columns
List of features with the highest difference in qvalues
| DAE | PI | frequency | Differential Analysis Comparison | diff_qvalue | |
|---|---|---|---|---|---|
| protein groups | |||||
| Q9Y6C2 | 0.028 | 0.995 | 119 | DAE (yes) - PI (no) | 0.967 |
| A0A087WXE9;E9PQ70;Q6UXH9;Q6UXH9-2;Q6UXH9-3 | 0.011 | 0.976 | 108 | DAE (yes) - PI (no) | 0.965 |
| P22692;P22692-2 | 0.035 | 0.987 | 170 | DAE (yes) - PI (no) | 0.952 |
| P48745 | 0.012 | 0.955 | 180 | DAE (yes) - PI (no) | 0.943 |
| P22748 | 0.033 | 0.976 | 159 | DAE (yes) - PI (no) | 0.943 |
| ... | ... | ... | ... | ... | ... |
| A0A0J9YXX1 | 0.047 | 0.057 | 197 | DAE (yes) - PI (no) | 0.010 |
| F5GY80;F5H7G1;P07358 | 0.046 | 0.057 | 197 | DAE (yes) - PI (no) | 0.010 |
| Q9NX62 | 0.045 | 0.055 | 197 | DAE (yes) - PI (no) | 0.010 |
| P00740;P00740-2 | 0.043 | 0.053 | 197 | DAE (yes) - PI (no) | 0.010 |
| K7ERG9;P00746 | 0.042 | 0.052 | 197 | DAE (yes) - PI (no) | 0.010 |
206 rows × 5 columns
Differences plotted with created annotations#
pimmslearn.plotting - INFO Saved Figures to runs/alzheimer_study/diff_analysis/AD/PI_vs_DAE/diff_analysis_comparision_1_DAE
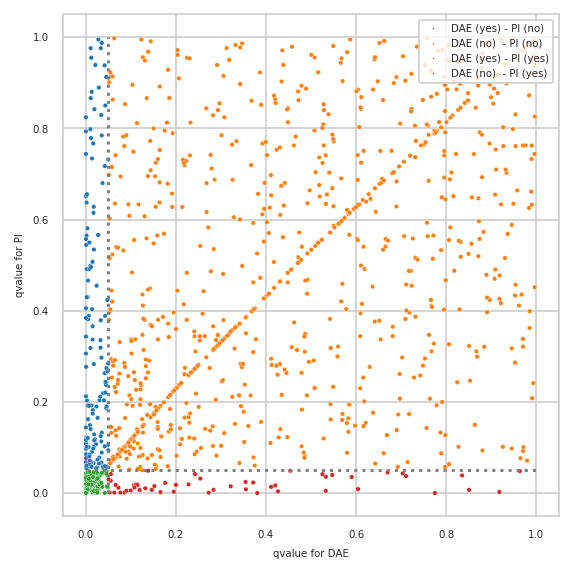
also showing how many features were measured (“observed”) by size of circle
pimmslearn.plotting - INFO Saved Figures to runs/alzheimer_study/diff_analysis/AD/PI_vs_DAE/diff_analysis_comparision_2_DAE
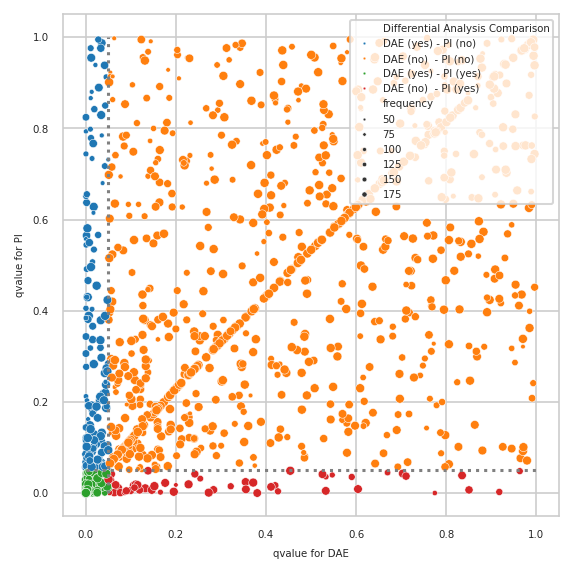
Only features contained in model#
this block exist due to a specific part in the ALD analysis of the paper
root - INFO No features only in new comparision model.
DISEASES DB lookup#
Query diseases database for gene associations with specified disease ontology id.
pimmslearn.databases.diseases - WARNING There are more associations available
| ENSP | score | |
|---|---|---|
| None | ||
| PSEN1 | ENSP00000326366 | 5.000 |
| APP | ENSP00000284981 | 5.000 |
| PSEN2 | ENSP00000355747 | 5.000 |
| APOE | ENSP00000252486 | 5.000 |
| TREM2 | ENSP00000362205 | 4.825 |
| ... | ... | ... |
| CEP170B | ENSP00000404151 | 0.683 |
| SMPDL3A | ENSP00000357425 | 0.683 |
| ADAMTS10 | ENSP00000471851 | 0.683 |
| PPP3R2 | ENSP00000498330 | 0.683 |
| VAT1 | ENSP00000347872 | 0.683 |
10000 rows × 2 columns
only by model#
Only by model which were significant#
Only significant by RSN#
mask = (scores_common[(str(args.baseline), 'rejected')] & mask_different)
mask.sum()